Biochemistry 2280A Lecture Notes - Lecture 11: Steric Effects, Coil Spring, Carboxylic Acid
Get access


Related Documents
Related Questions
1. A sequence of amino acids in a certain protein is found to be â Ser â Gly â Pro â Gly â
The sequence is most probably part of a(n):
a. beta turn
b. antiparallel beta sheet
c. alpha-helix
d. parallel beta-sheet
e. alapha-sheet
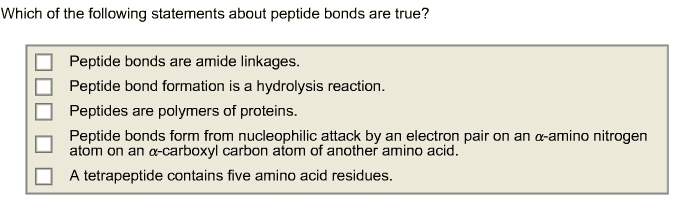
2. Which statement about peptides bonds is NOT TRUE?
| There is perfectly free rotation about the peptide bond. | |||||||||||
| The peptide bond has partial double-bond character. | |||||||||||
| The peptide bond is shorter than a normal carbon-nitrogen single bond. | |||||||||||
| The peptide bond usually has a trans configuration with respect to the alpha-carbons of the two amino acids involved in the peptide bond. | |||||||||||
| Peptide bonds are made via condensation of an amine and a carboxylic acid with loss of water. 3. Which of the following statements about proteins is FALSE?
| |||||||||||